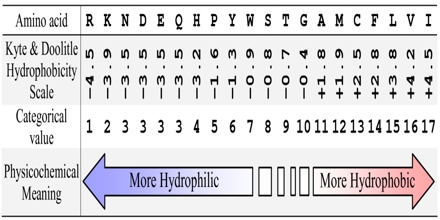
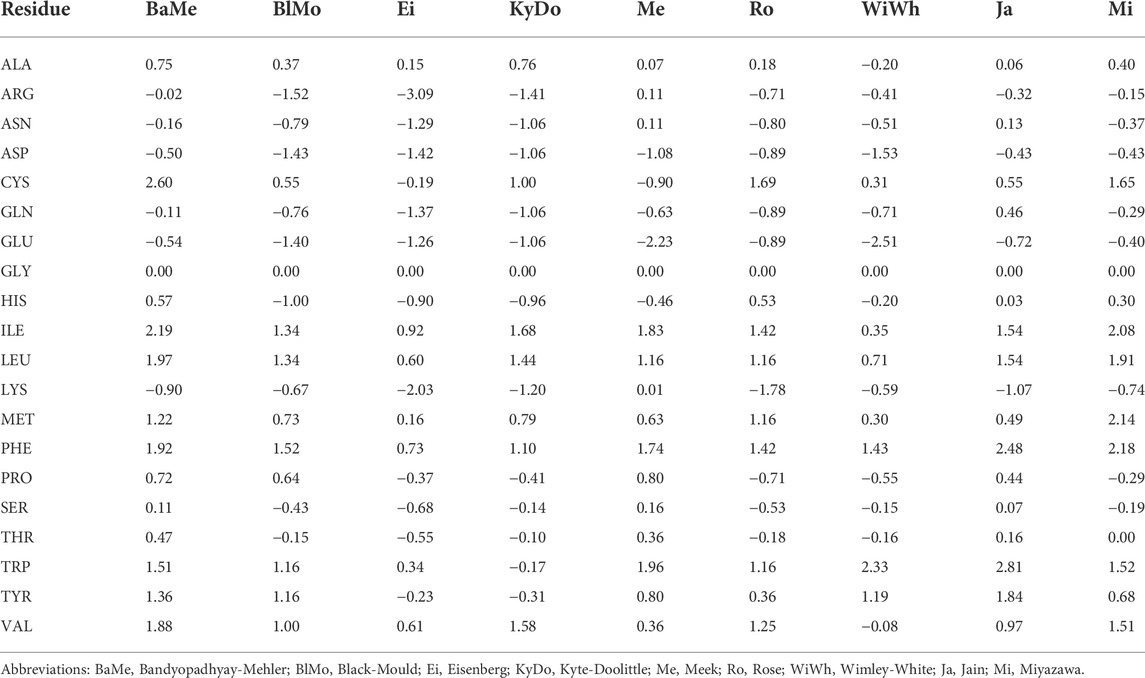
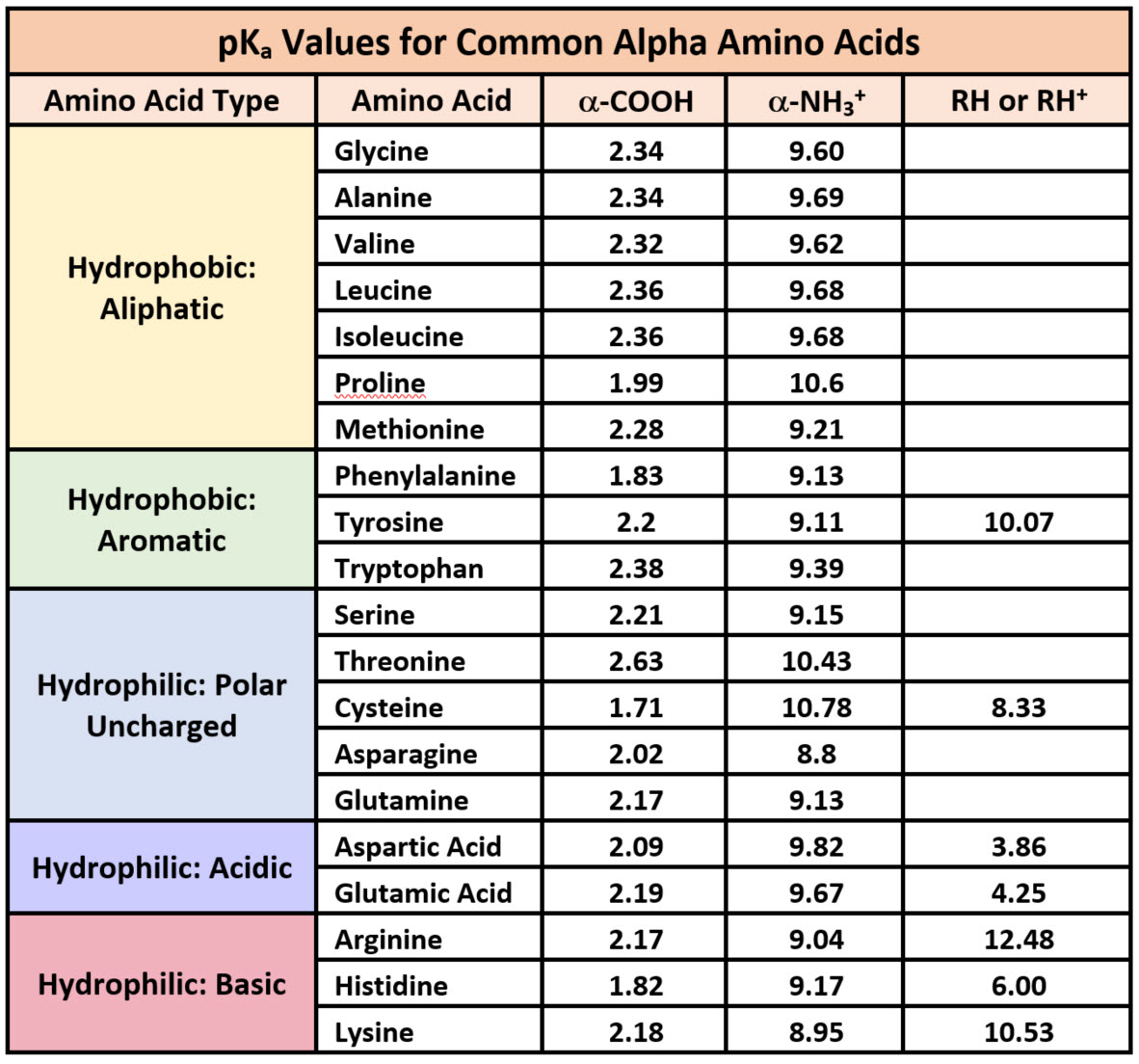
50 years of amino acid hydrophobicity scales: revisiting the capacity for peptide classification | Biological Research | Full Text
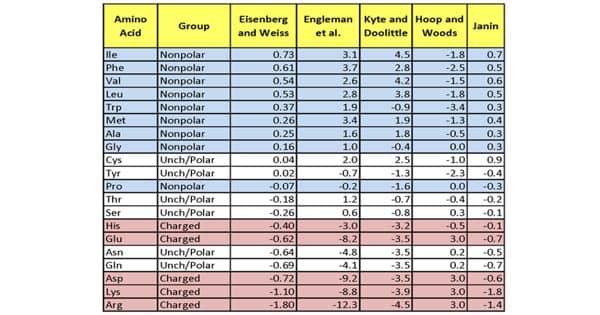
Hydrophobicity Scales - more Hydrophobic Amino Acids have Highest Positive Values - Assignment Point
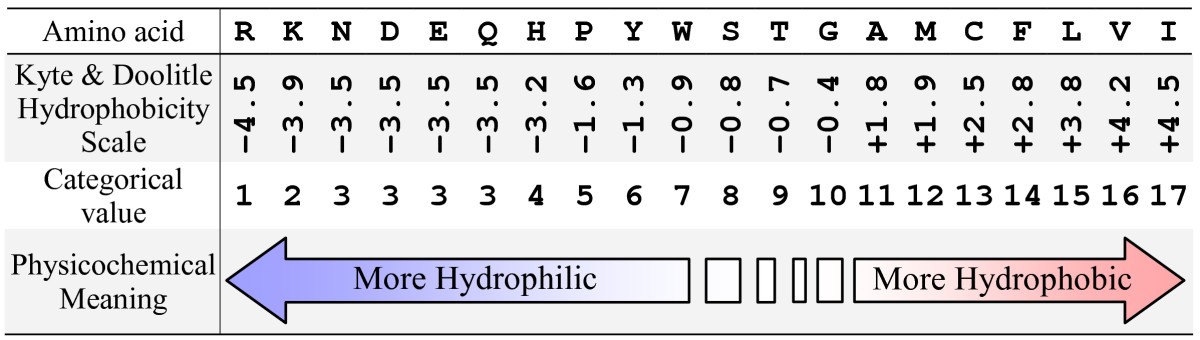
Searching for interpretable rules for disease mutations: a simulated annealing bump hunting strategy | BMC Bioinformatics | Full Text

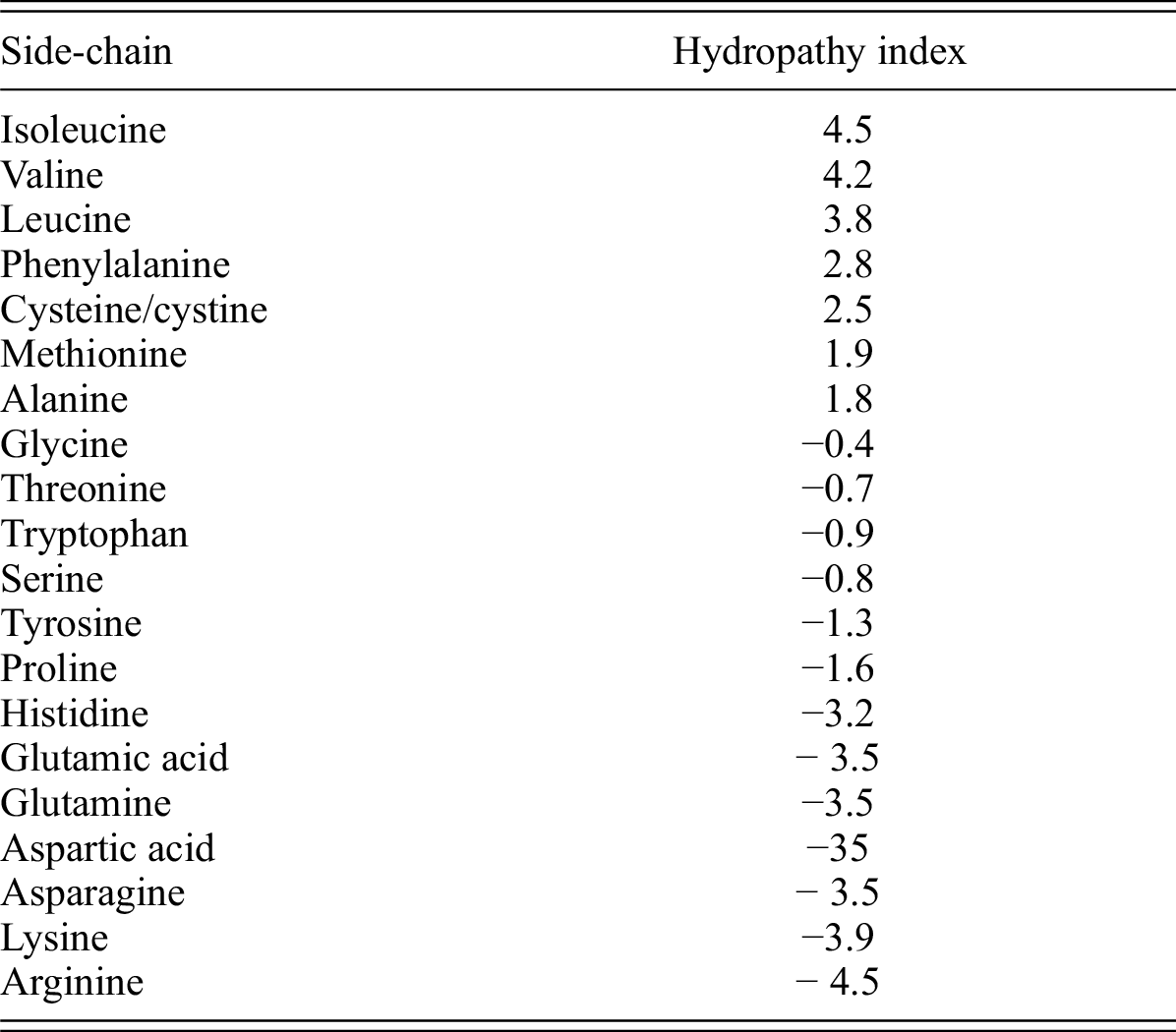
Side-chain hydrophobicity scale derived from transmembrane protein folding into lipid bilayers | PNAS
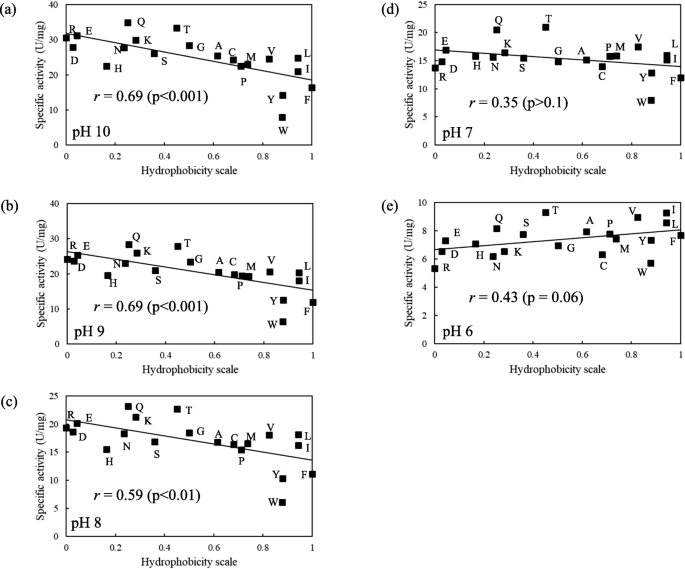
The hydrophobicity of an amino acid residue in a flexible loop of KP-43 protease alters activity toward a macromolecule substrate | Applied Microbiology and Biotechnology

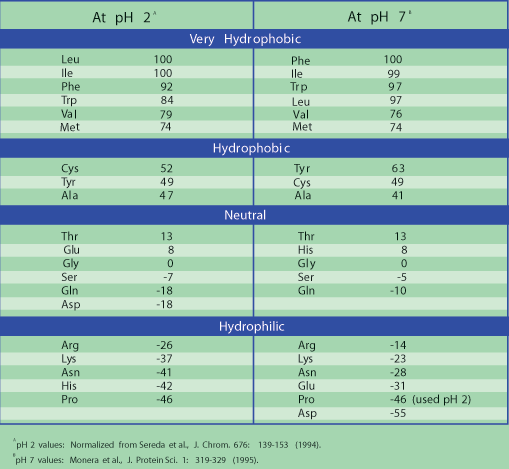
N-Capping Motifs Promote Interaction of Amphipathic Helical Peptides with Hydrophobic Surfaces and Drastically Alter Hydrophobicity Values of Individual Amino Acids | Analytical Chemistry



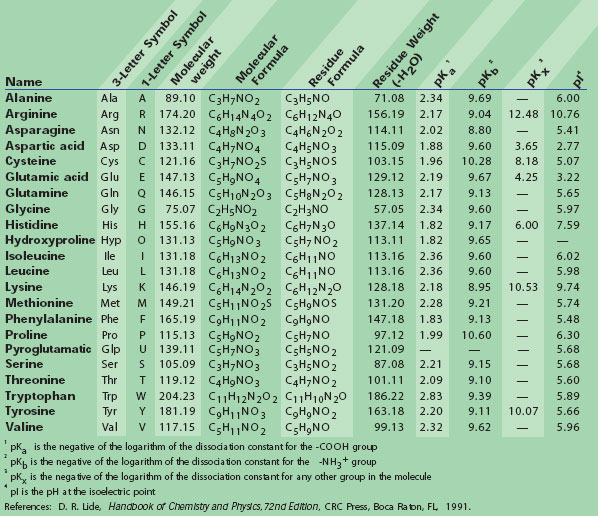

![PDF] A hydrophobic proclivity index for protein alignments | Semantic Scholar PDF] A hydrophobic proclivity index for protein alignments | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f46db37dac679e2f3424b0f388354cb97f9ac808/5-Table1-1.png)